Proteins structures are complicated (really complicated), and the code we use to describe their sequences needs to reflect this by being concise and easy to understand.
As scientists became able to identify and synthesise longer and longer sequences of amino acids, the notation they used to characterise them had to adapt, evolve, and (crucially) condense. As a snapshot of this process beginning, in 1904 it was evidently the height of condensed terminology was to give dipeptides easy names: Fischer explains how his newly coined name ‘proline’ could be appended to ‘alanine’ to describe the dipeptide, ‘prolylalanine’.1
As things moved on, and longer peptides needed to be described, three-letter notation was created. These were generally self-explanatory (e.g. alanine is Ala, etc.), and could be strung together to describe the sequence. This sufficed for a while, but in 1958 the possibility of a more condensed, one-letter code was first raised.2
A few one-letter systems were suggested and trialled over the following decade, but the system we know today was formally established in a paper published by the IUPAC-IUB Commission on Biochemical Nomenclature (CBN) in 1968.2 Their “tentative rules” were based on a combination of the systems previously suggested, and the authors provide (mostly) useful explanations for the theory behind the choices of letters, which I will proceed to work through here:
One-Letter Allocations
The one-letter system was designed to accommodate the twenty common amino acids, for which the 26 letters of the alphabet were, thankfully, sufficient. In fact, there were letters to spare, and three were removed from the final selection: U was eliminated as it can be confused with V in handwritten materials; O was disregarded as it could be confused with zero or, in low-quality printing, with C, D, G or Q; and J was disqualified for “linguistic reasons”, in that it is absent in several languages.

Where there was no ambiguity, the first letter of the amino acid name was assigned. This was possible in six cases: cysteine, histidine, isoleucine, methionine, serine, and valine.
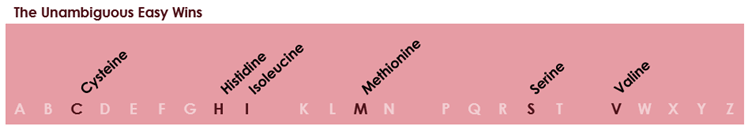
After this, things got more complicated, and increasingly arbitrary. The rest of the amino acids all shared a first letter with at least one other example, so a decision had to be made as to which would get to take the easy option.

The selections were made based on which compounds were the most frequently occurring or structurally simple: alanine, glycine, leucine, proline and threonine.

A level of logic was applied to the next allocations. Two amino acids were described as being “phonetically suggestive”: F for phenylalanine and R for arginine. Then, the double-ring structure of tryptophan was associated with the “bulky letter W” (i.e. double ring – double U).
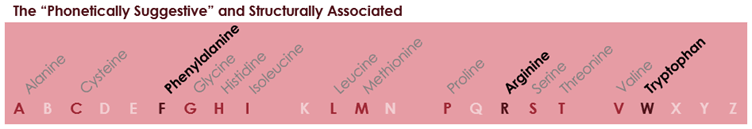
The explanation provided in the article then entirely breaks down:
“The letters N and Q are assigned to asparagine and glutamine, respectively; D and E are assigned to aspartic acid and glutamic acid, respectively”
Helpful!

While there doesn’t appear to be much logic here, there are some clues to the thought process if you go a bit further back in the literature, and apply a little bit of inference and guesswork. Firstly, there just aren’t many letters left to choose from! So without an obvious answer, any vaguely reasonable suggestion was likely to stick unless disputed. A previous set of suggestions was published by Šorm et al. in 1961, which gives some hints. They provide their own suggestions alongside those of a contemporary, Hans Neurath, with whom they had discussed the matter. Neurath had devised a system based on using vowels for acidic and basic residues, and consonants for everything else. The table from the manuscript is reproduced here:3

We can see quite a few differences between this table and the code we use today. However, there are a couple of things that jump out. Firstly, the assignment of N for asparagine matches our current system. The authors have put the n in the full name in bold, presumably by way of explanation (as if there isn’t an n in all but 2 of the other names), but they don’t deign to comment on this further in the text. Secondly, there is some familiarity with glutamic acid and glutamine as well. Neurath uses E for both of these, in the same way that he suggests A for both aspartic acid and asparagine. This is presumably related to the fact that for a long time proteins were broken up into amino acids by harsh hydrolysis which hydrolysed the amides to the carboxylic acids, and you couldn’t tell the difference between the two anyway. So we could generously assume that Neurath’s E for glutamic acid stuck, and perhaps D was later selected for aspartic acid simply by being the previous letter in the alphabet.
Šorm’s suggestion of Q for glutamine comes with the delightfully non-sensical explanation:
“…in many other cases characteristic symbols derived either from the spelling (ala- l, arg-r, lys-I, leu- u, ileu- w, phe- f) or from the graphical representation (glu- g, glu(NH2)- q) were chosen.”
I struggle to see the logical connection between “glu(NH2)” and the letter q, unless we are supposed to somehow associate the subscript number with the tail of the q. However, it is possible that I am failing to understand what an amino acid “graphical representation” meant in the 1960s.
Later descriptions of these assignments add a little bit more logic onto them, but we can’t assume that this isn’t post-rationalisation. The 1969 edition of the Atlas of Protein Sequence and Structure,4 which references the “tentative rules” published in 1968, notes that the smaller molecules (aspartic acid and asparagine) come before the larger ones (glutamic acid and glutamine) in the alphabet.*
The final two amino acids, lysine and tyrosine, are said to have been assigned the letters that were closest alphabetically from the ones left. If you ignore X, this works fine (more on X below); some possible post-rationalisation from 1969 includes tyrosine in the “phonetically suggestive” category due to the second letter being Y.4*

Finally, we have the added extras. X was assigned to situations where the amino acid is unknown, or is atypical, which fits with how we generally use this letter. Then B and Z were assigned as semi-wild cards for situations when the difference between glutamic acid and glutamine (Z), or aspartic acid and asparagine (B) can’t be determined, for reasons explained previously. This is clearly a case of using up the last remaining letters, but it is notable that the order remains the same, with the smaller asp- molecules coming first alphabetically.

This core code, put forward as “tentative rules” in 1968 has remained unchanged now for half a century. Some of them may be difficult to remember, but the fact that there was at least some logic behind the assignments may be helpful…or not.
There have been a few minor additions to this code since, for special cases.5 The rare amino acids pyrrolysine and selenocysteine are sometimes assigned the letters O and U, respectively. Both of these letters were originally discarded for legibility issues, but that hardly seems relevant nowadays. The letter J is also sometimes used to mean either isoleucine or leucine in situations when they can’t be distinguished. All of which is excellent news for people whose names start with J and O who happen to want to draw out their peptide chain signatures!

References
1. Fischer E, Suzuki U. Synthese von Polypeptiden. III. Derivate der α-Pyrrolidincarbonsaure. Berichte der Dtsch chernischen Gesellschaft. 1904;37:2842-2848.
2. Hoffmann-Ostenhof O, Cohn WE, Braunstein AE, et al. IUPAC-IUB Commission on Biochemical Nomenclature. A One-Letter Notation for Amino Acid Sequences. J Biol Chem. 1968;243(13):3557-3559. doi:10.1021/bi00848a001. URL:https://www.jbc.org/content/243/13/3557.long
3. Šorm F, Keil B, Vaněček J, et al. On proteins. LXIII. Lower structures in the chains of proteins. Collect Czechoslov Chem Commun. 1961;26(2):531-578. doi:10.1135/cccc19610531. URL:http://gen.lib.rus.ec/scimag/10.1135%2Fcccc19610531
4. Dayhoff MO. Atlas of Protein Sequence and Structure. 4th ed.; 1969. https://play.google.com/books/reader?id=6_AG_Q8cXIwC&hl=en_GB&pg=GBS.RA4-PA2URL:https://play.google.com/books/reader?id=6_AG_Q8cXIwC&hl=en_GB&pg=GBS.RA4-PA2
5. Wikipedia. https://en.wikipedia.org/wiki/FASTA_formatURL:https://en.wikipedia.org/wiki/FASTA_format
*It is possible that the explanations I am suggesting are post-rationalisations in the 1969 edition of the Atlas of Protein Sequence and Structure were already present in the 1965 or 1967–8 editions and contributed towards the rules put forward by the committee. However, these versions are not available as ebooks and at time of writing going to find these in a library isn’t really an option.
Awesome ! This is a rare knowledge and hardly any textbook mentions these things. Thanks from India.
LikeLike
I guess the “struggle to see the logical connection between “glu(NH2)” and the letter q” in the case of (glu- g, glu(NH2)- q) is because you don’t pay atention to the shape of both letters, not to the chemical compound. My opinion is that letter “g” and “q” are similiar in a way that “G” and “Q” are not. Like capital “L” and lowercase “i” comparing to lowercase “l” and capital “I”, E and 3, 5 and S, T and 7, H and N, n and m, C and G, Z and S, D and O, K and X, UVY, WM, AV, un, nm, mw, rL, JL, bdpq…
LikeLike